Dortmund, 11th November 2022
Facebook founder Mark Zuckerberg and his wife Priscilla Chan support two research projects at ISAS with 40,000 US dollars. Over the course of one year, the Chan Zuckerberg Initiative (CZI) will fund the development of the Dortmund software for the napari image analysis platform: At ISAS, the research groups AMBIOM* - Analysis of Microscopic BIOMedical Images and Spatial Metabolomics* will develop new plug-ins. Thus, scientists worldwide will be able to better analyze microscopic and chemical images – free of charge.
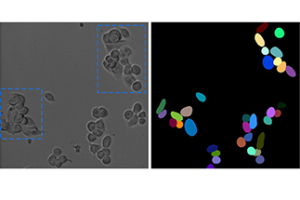
The left picture shows a microscopic image of tumor cells. In the right view, segmentation by means of common computer programs can be seen. As soon as the cells are close to each other or overlap (see blue boxes in the first image), segmentation deteriorates. As a result, fully automated tracking leads to inaccuracies.
© Prof. Dr. Barbara Grüner / Universitätsklinikum Essen
Cell movements provide important insights in biomedicine, for example regarding the development of various forms of cancer. To find out how tumor cells spread in the body in different types of cancer, researchers among other things examine their movement under the microscope. The resulting time-lapse recordings provide information, for example, about the motility of the cells. For this quantitative analysis, cells that overlap must first be separated from one another (segmentation). Then, their paths can be tracked.
Artificial intelligence improves cell tracking
Image analysis has improved significantly in recent years. Nevertheless, the technology currently still reaches its limits when it comes to cell tracking. "Right now, there are many microscopy scenarios in which automated analysis does not provide satisfactory results - and manual curation is required. The images of 50 cells generate large amounts of data which researchers can hardly analyze manually," explains Dr Jianxu Chen, an artificial intelligence (AI) expert and head of AMBIOM. That is why the computer scientist and his team want to develop specific software in the napari ecosystem. This software will enable biomedical scientists to intervene directly in the automatic analysis and correct errors, for example during segmentation or tracking.
The "Human-in-the-loop Cell Tracking" software, which has received funding of 20,000 US dollars, will include a total of three modules (segmentation, tracking and analysis). Its goal using AI: first, to improve the result of napari image analysis, and second, to train the algorithm with the human-input information using machine learning. To this end, the AMBIOM team will cooperate with immunologists at ISAS, University Hospital Essen and the University of Duisburg-Essen.
Mass spectrometry image data for worldwide exchange
AMBIOM is also involved in the programming of the second software for biochemical imaging, which is funded with another 20,000 US dollars. This project is the first napari plug-in for the evaluation and annotation of mass spectrometry imaging data. A mass spectrometer can be used to identify substances such as metabolites in a sample based on their masses. Using mass spectrometry imaging (MSI), scientists can, for example, examine tumor tissue for metabolic differences at subcellular resolution. In this way, they obtain information about the spatial distribution of the molecules and can compare the results with morphological abnormalities in the tissue.
Dr Prasad Phapale, a chemist and head of Spatial Metabolomics, is striving to improve the multiplexing (integration) of MSI data with other image formats. For example, the "Biochemical Annotations of Mass Spectrometry Imaging Data" plug-in for napari will allow biochemical annotation of MSI data with image co-registration (image fusion). In short, it will enable scientists worldwide to match their MSI results with metabolite databases, as well as match spatial information from their imaging with that of complementary analytical methods, such as microscopy. This would improve the sharing of knowledge within the research community.
About napari
napari is an open source tool that enables a powerful visualization of multidimensional images, for example from microscopy. napari is based on the Python programming language. It is being continuously expanded with the help of a worldwide growing community of researchers and software developers. CZI promotes napari through its Imaging Program with the goal of facilitating biologists' access to new methods of image analysis based on machine learning.
* The MSCoreSys associated junior research group AMBIOM – Analysis of Microscopic BiOMedical Images is funded by the Federal Ministry of Education and Research (funding reference 161L0272). The Federal Ministry for Education and Research is also funding the MSCoreSys-associated junior research group Spatial Metabolomics (funding number 161L0271).